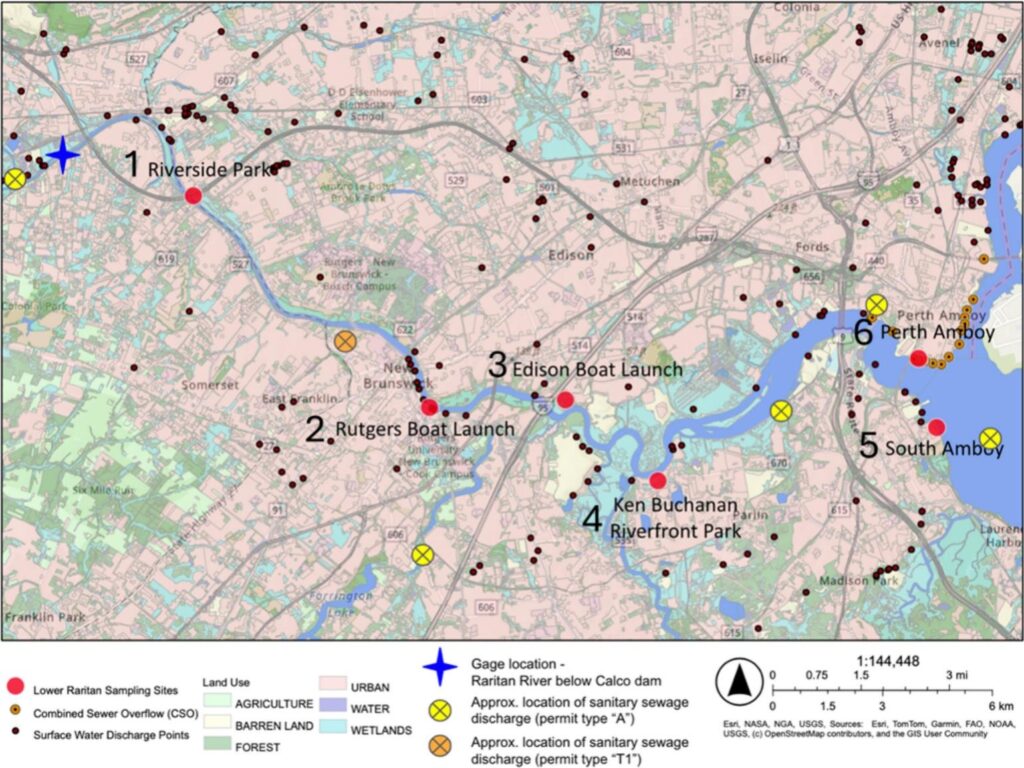
Viability-based DNA techniques show promise for fecal pollution source tracking in NJ Sea Grant funded study
With huge thanks to our partners including Rutgers Cooperative Extension of Middlesex County, the Fahrenfeld Lab at Rutgers – The State University of New Jersey, and NJ Sea Grant, the LRWP is pleased to share that findings based on our Summer Raritan River civic science pathogens monitoring and data gathering efforts were published by Elsevier in Science of the Total Environment:
Ehasz, G., Almosd, L., Ahamed, P., Bakacs, M., Fenyk, H., Fahrenfeld, N.L. 2026. Comparison of propidium monoazide and total-DNA based qPCR and long-read sequencing for microbial source tracking in an estuarine river. Science of the Total Environment. 1013, 181271. 10.1016/j.scitotenv.2025.181271
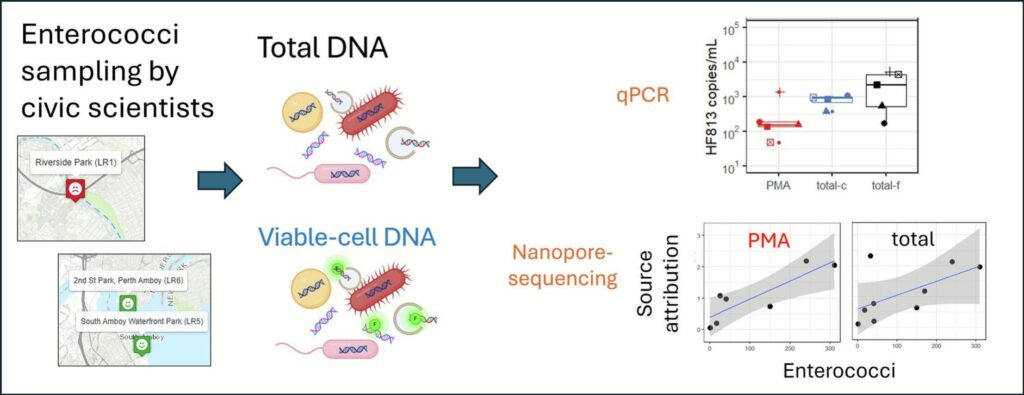
Recap: Fecal pollution is a common cause of water pollution and improved microbial source tracking techniques are needed. A NJ Sea Grant funded team from Rutgers, Middlesex County Extension, and the Lower Raritan Watershed Partnership conducted microbial monitoring on the Lower Raritan River. Total-DNA and viable-cell DNA techniques were compared.
Results: Using viable-cell DNA rather than total DNA resulted in better correlations between fecal pollution and source measurements. The techniques demonstrated in the Lower Raritan can be applied in any watershed with fecal pollution issues and may improve results.